FGFR1 FISH Probe
Our FGFR1 probe is designed to detect FGFR1 amplifications and deletions. The probe comes labeled in orange, but can be customized to meet your needs.
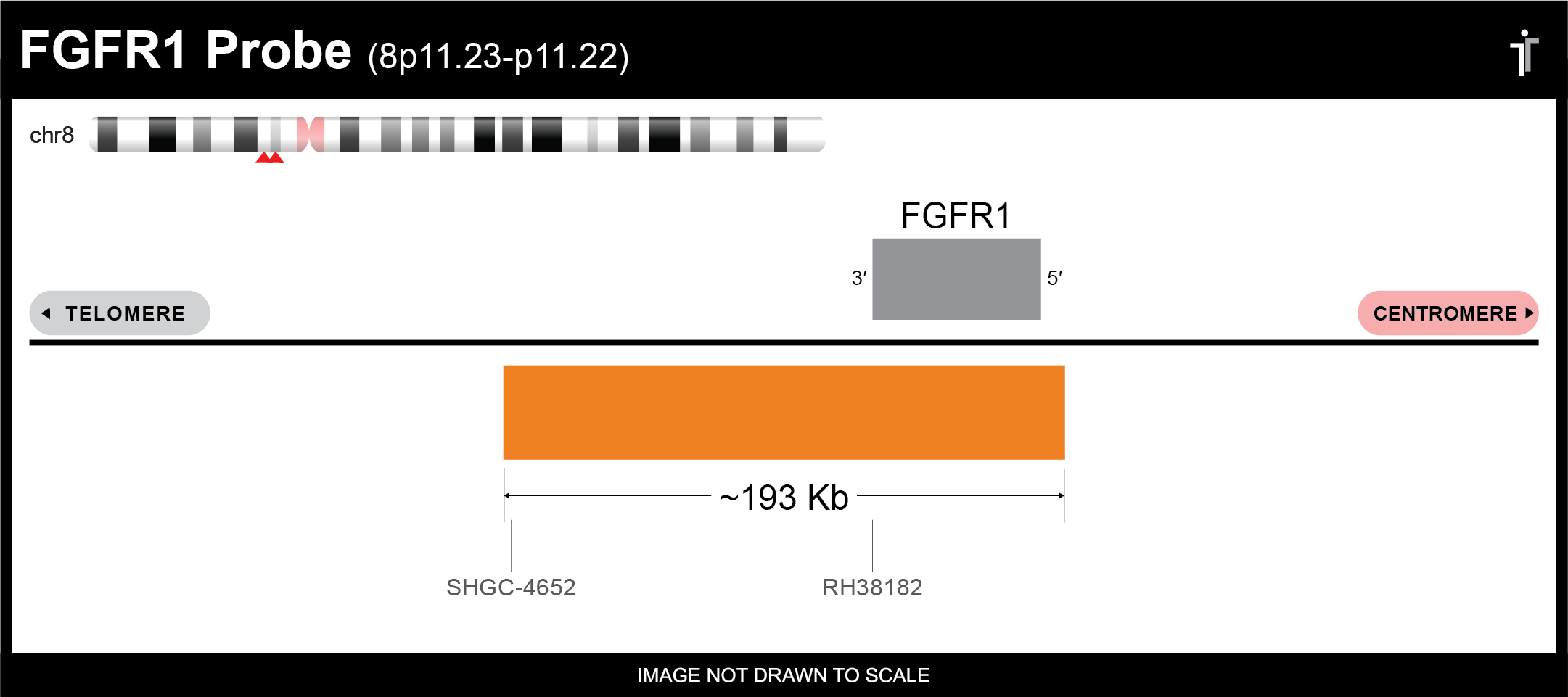
** This product is for in vitro and research use only. This product is not intended for diagnostic use.
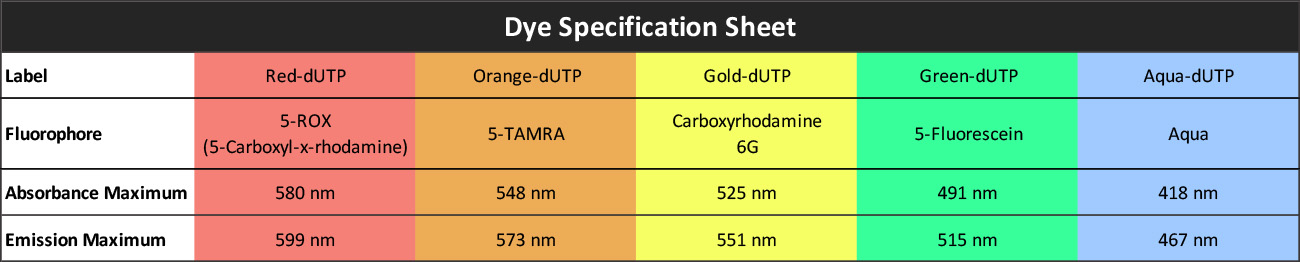
| SKU | Test Kits | Buffer | Dye Color | Order Now |
|---|---|---|---|---|
| FGFR1-20-OR (Standard Design) | 20 (40 μL) | 200 μL |

|
|
| FGFR1-20-RE | 20 (40 μL) | 200 μL |

|
|
| FGFR1-20-AQ | 20 (40 μL) | 200 μL |

|
|
| FGFR1-20-GR | 20 (40 μL) | 200 μL |

|
|
| FGFR1-20-GO | 20 (40 μL) | 200 μL |

|
Gene Summary
The protein encoded by this gene is a member of the fibroblast growth factor receptor (FGFR) family, where amino acid sequence is highly conserved between members and throughout evolution. FGFR family members differ from one another in their ligand affinities and tissue distribution. A full-length representative protein consists of an extracellular region, composed of three immunoglobulin-like domains, a single hydrophobic membrane-spanning segment and a cytoplasmic tyrosine kinase domain. The extracellular portion of the protein interacts with fibroblast growth factors, setting in motion a cascade of downstream signals, ultimately influencing mitogenesis and differentiation. This particular family member binds both acidic and basic fibroblast growth factors and is involved in limb induction. Mutations in this gene have been associated with Pfeiffer syndrome, Jackson-Weiss syndrome, Antley-Bixler syndrome, osteoglophonic dysplasia, and autosomal dominant Kallmann syndrome 2. Chromosomal aberrations involving this gene are associated with stem cell myeloproliferative disorder and stem cell leukemia lymphoma syndrome. Alternatively spliced variants which encode different protein isoforms have been described; however, not all variants have been fully characterized. [provided by RefSeq, Jul 2008]
Gene Details
Gene Symbol: FGFR1
Gene Name: Fibroblast Growth Factor Receptor 1
Chromosome: CHR8: 38268655-38326352
Locus: 8p11.23
FISH Probe Protocols
| Protocol, Procedure, or Form Name | Last Modified | Download |
|---|